Whipworm microbiome study: Patent human infections with the whipworm, Trichuris trichiura, are not associated with alterations in the faecal microbiota
Cooper P, Walker AW, Reyes J, Chico M, Salter SJ, et al. (2013) PLoS ONE 8(10): e76573.
Project Summary
Background: The soil-transmitted helminth (STH), Trichuris trichiura colonises the human large intestine where it may modify inflammatory responses, an effect possibly mediated through alterations in the intestinal microbiota. It was hypothesised that patent T. trichiura infections would be associated with altered faecal microbiota and that anthelmintic treatment would induce a microbiota resembling more closely that observed in uninfected individuals.
Materials and methods: School children in Ecuador were screened for STH infections and allocated to 3 groups: uninfected, T. trichiura only, and mixed infections with T. trichiura and Ascaris lumbricoides. A sample of uninfected children and those with T. trichiura infections only were given anthelmintic treatment. Bacterial community profiles in faecal samples were studied by 454 pyrosequencing of 16 S rRNA genes.
Results: Microbiota analyses of faeces were done for 97 children: 30 were uninfected, 17 were infected with T. trichiura, and 50 with T. trichiura and A. lumbricoides. Post-treatment samples were analyzed for 14 children initially infected with T. trichiura alone and for 21 uninfected children. Treatment resulted in 100% cure of STH infections. Comparisons of the microbiota at different taxonomic levels showed no statistically significant differences in composition between uninfected children and those with T. trichiura infections. There was found to be a decreased proportional abundance of a few bacterial genera from the Clostridia class of Firmicutes and a reduced bacterial diversity among children with mixed infections compared to the other two groups, indicating a possible specific effect of A. lumbricoides infection. Anthelmintic treatment of children with T. trichiura did not alter faecal microbiota composition.
Discussion: This data indicate that patent human infections with T. trichiura may have no effect on faecal microbiota but that A. lumbricoides colonisation might be associated with a disturbed microbiota. Our results also catalogue the microbiota of rural Ecuadorians and indicate differences with individuals from more urban industrialised societies.
Resources
Full manuscript is available at:
Patent Human Infections with the Whipworm, Trichuris trichiura, Are Not Associated with Alterations in the Faecal Microbiota
Philip Cooper (equal contributor),Alan W. Walker (equal contributor),Jorge Reyes,Martha Chico,Susannah J. Salter,Maritza Vaca,Julian Parkhill. Oct. 4th, 2013 PLoS One DOI: 10.1371/journal.pone.0076573
All raw 454 sequence data is available in the European Nucleotide Archive (ENA) short-read archive under the sample accession number ERS236495 and/or the study accession number ERP002465.
Data table showing Trichuris trichiura effect on microbiomes in Ecuador:
Ecuador.xlsx
Supporting Table S3, containing all OTU distribution and taxonomic classification data, can be downloaded here:
Whipworm_OTU.xlsx
Supporting Table S4, containing the representative sequences for all OTUs, can be downloaded here:
Whipworm_OTU_sequences.xlsx
Selected Results from the Study
Table 1: Characteristics of study population stratified by infection status with soil-transmitted helminth infections. Epg = eggs per gram.

Table 2: Relative composition of faecal microbiota by bacterial genus in children with no STH infection (uninfected), children infected with only T. trichiura, and those infected with both T. trichiura and A. lumbricoides (mixed infection).
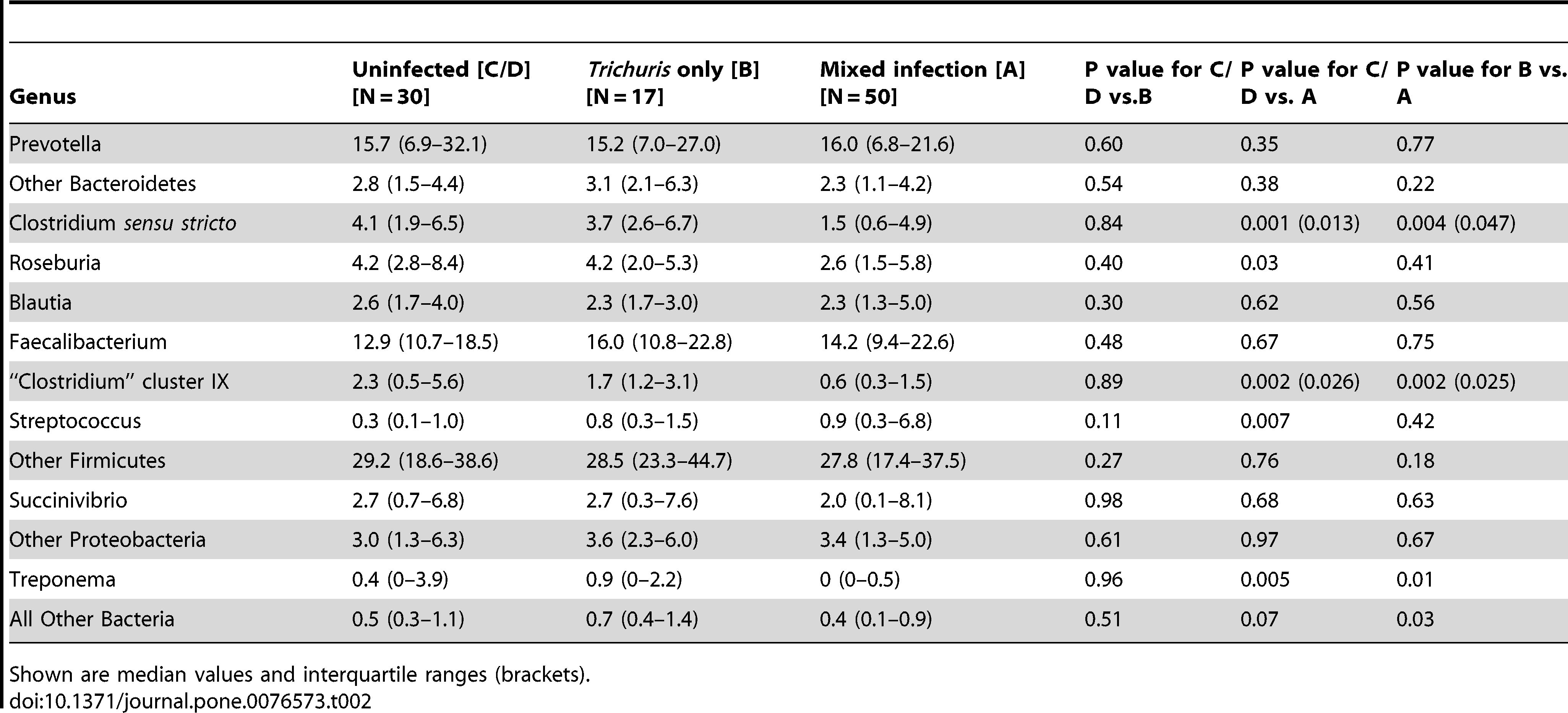
Table 3: Relative composition of faecal microbiota by bacterial genus before and after treatment for children infected with T. trichiura only, uninfected children, and all children that received anthelmintic treatment (all matched pairs).
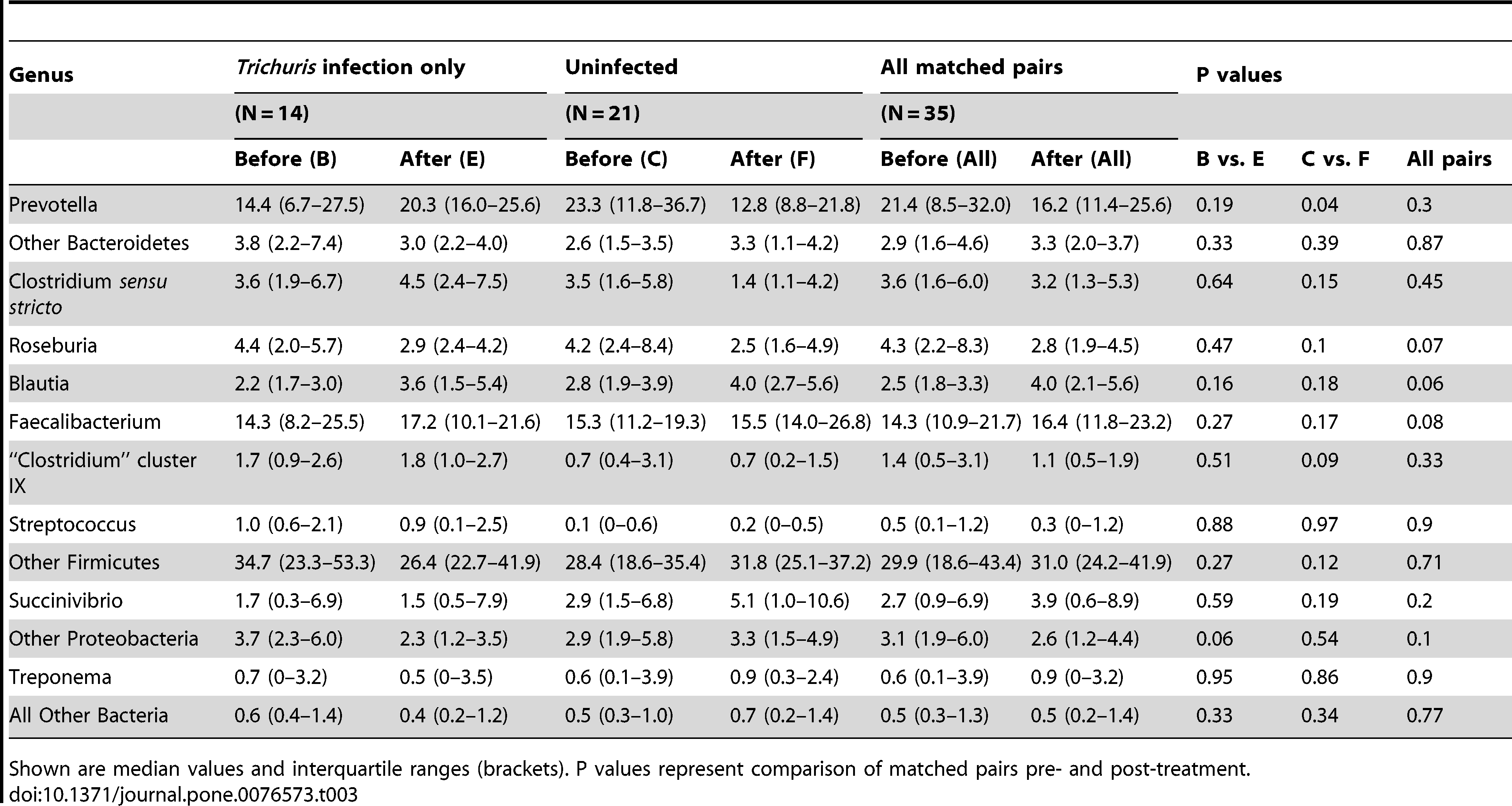
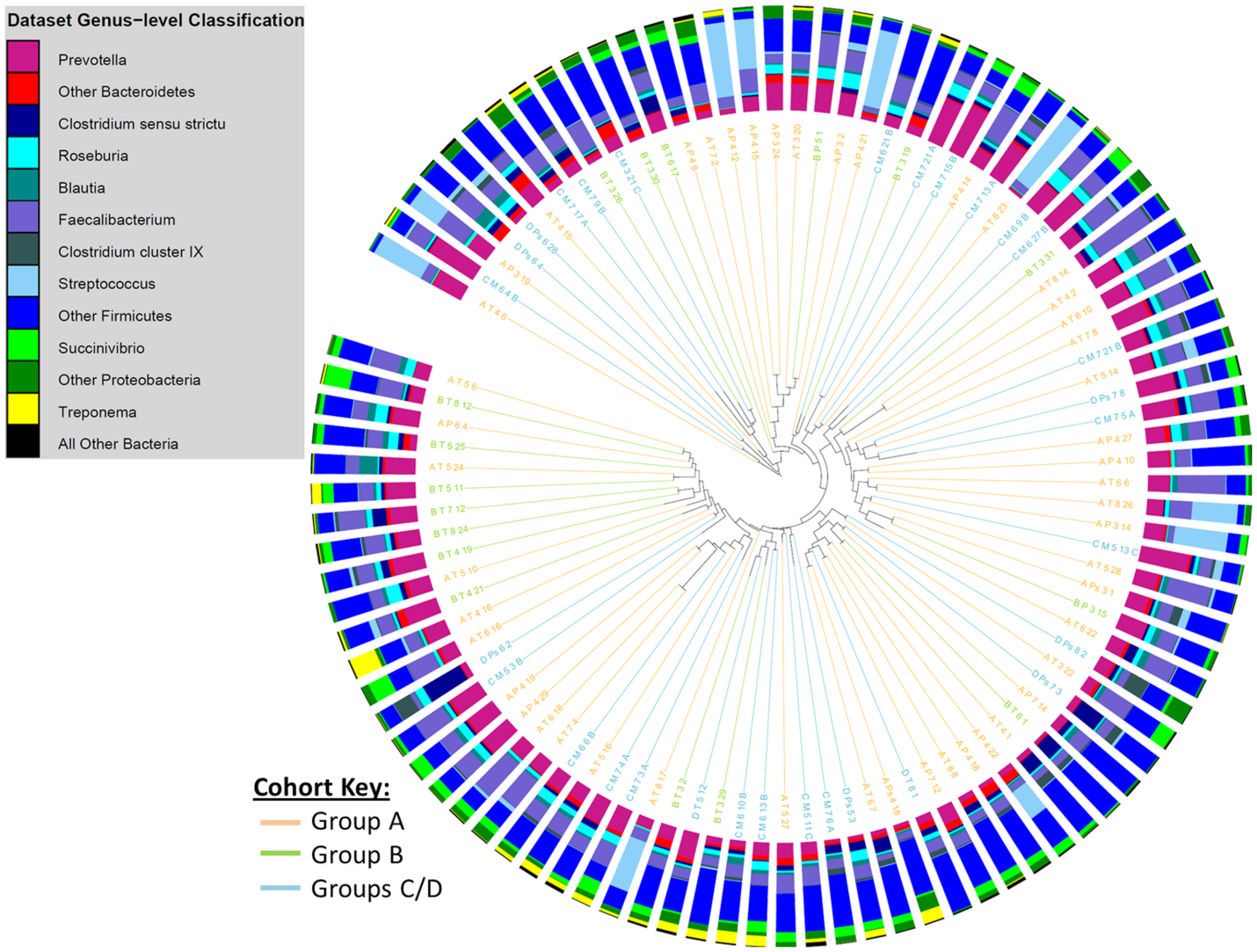
Figure 3: Cluster dendrogram, generated with the Jaccard calculator in the mothur software package, showing similarity in community membership at the OTU-level between faecal samples from different study groups.




