HelmCoP gene search - Build gene based custom queries against the HelmCoP database
|
How to use this page
Search the HelmCoP database for genes using supported queries. For information regarding gene naming for specific species, please refer to the HelmCoP FAQ. A typical HelmCoP user will first build a list of genes for the organism(s) they are interested in using the various filters provided. Genes of interest can be further explored by identifying the ortholog in which a given gene resides, and copy-pasting that ortholog name into a search constructed on the HelmCoP orthologs page. In this manner related genes meeting your filters can be found. HelmCoP can be searched using GO annotations, KEGG orthology (KO), and Interpro domains. Essentiality can also be used in the search to find genes that have RNAi phenotypes based on association with C. elegans genes. Genes with a PDB structure, signal peptide and a particular DrugBank ID can also be found. Further, HelmCoP can be searched for genes that are potential vaccine candidates. HelmCoP also provides extensive annotation of included proteins (see the chart at the bottom of this page). Output is originally displayed in HTML format, but a tab-delimited version of your query output is available via the Full Results Download button at the top of the results page. Please refer to the HelmCoP FAQ for additional information.
Due to the size of the database and the complexity of some queries, HelmCoP limits HTML output to the first 20000 rows returned by your query. Full results are only available via the Full Results Download button in the case of queries resulting in more than 20000 rows. Some extreme queries may be refused to prevent a page time-out, but in this case you will be prompted with the most 'expensive' portion of your query. In many cases simply removing the specified 'expensive' filter can result in a manageable output
|
Gene
|
Species
*Note that non-worm species are only annotated with drugbank, PPI, orthology assignments and PDB information
|
|
|
|
|
|
Vaccine candidates
|
|
|
Data availability notice
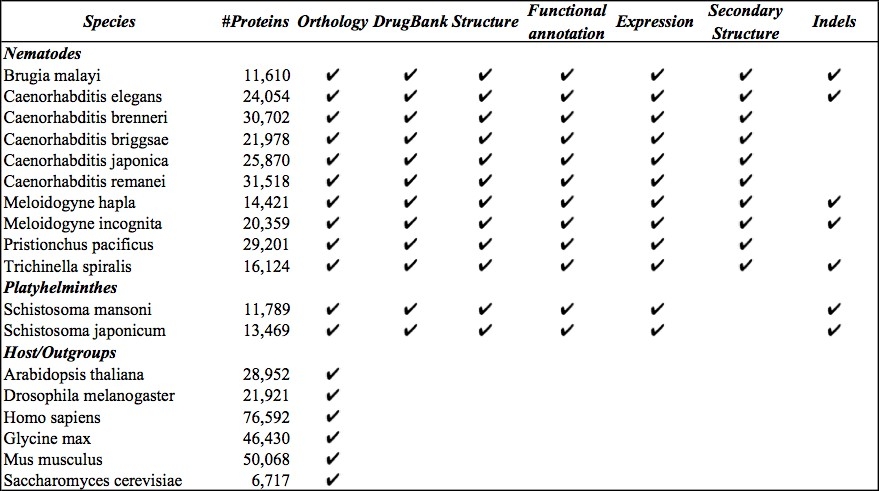
Not all analysis annotations are available for all organisms. Selecting filters or outputs that don't exist for your chosen organism(s) can produce empty result sets. Please refer to the table below to see which species have which annotations avaible to query.
|
|
|
|




